David A. Gutman
Department of Neurology, Emory University School of Medicine, Atlanta, GA, USA
MITI Minimum Information guidelines for highly multiplexed tissue images
Aug 21, 2021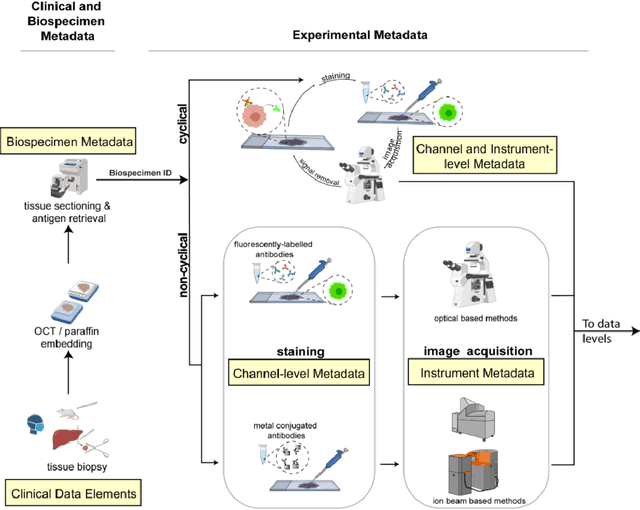
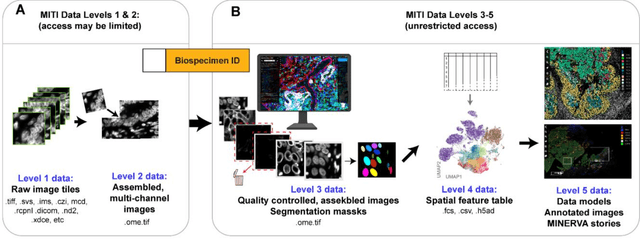
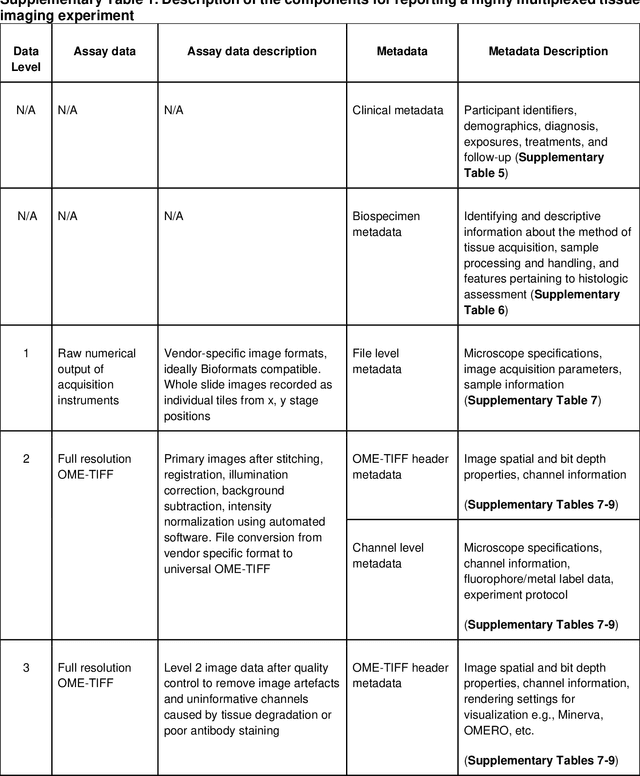
Abstract:The imminent release of atlases combining highly multiplexed tissue imaging with single cell sequencing and other omics data from human tissues and tumors creates an urgent need for data and metadata standards compliant with emerging and traditional approaches to histology. We describe the development of a Minimum Information about highly multiplexed Tissue Imaging (MITI) standard that draws on best practices from genomics and microscopy of cultured cells and model organisms.
NuCLS: A scalable crowdsourcing, deep learning approach and dataset for nucleus classification, localization and segmentation
Feb 18, 2021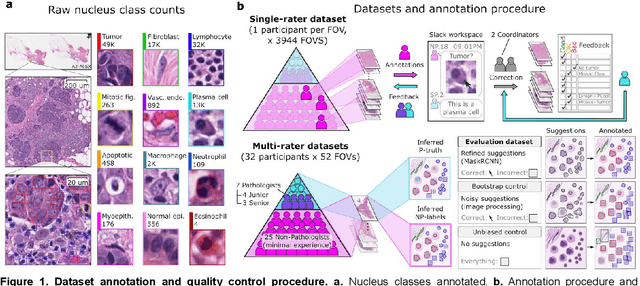
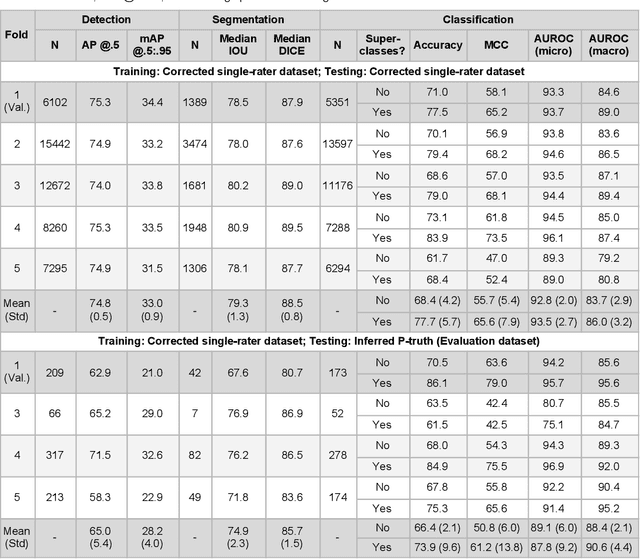
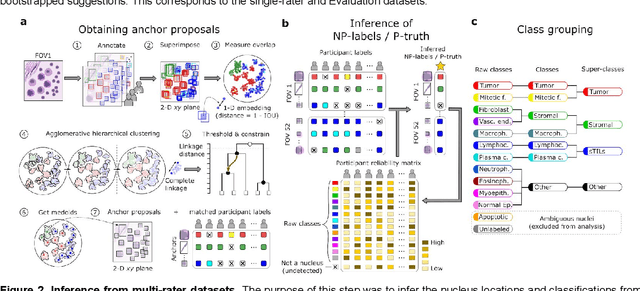
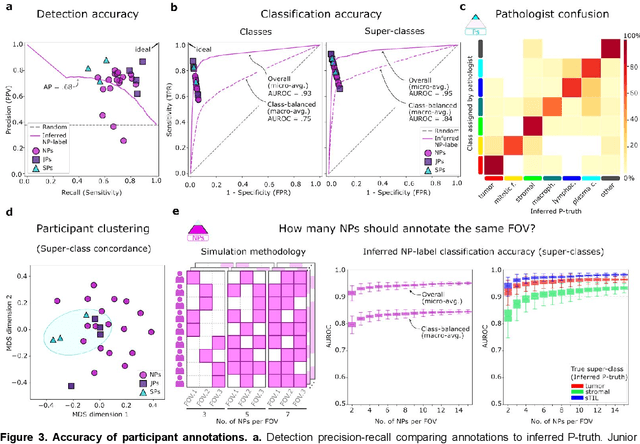
Abstract:High-resolution mapping of cells and tissue structures provides a foundation for developing interpretable machine-learning models for computational pathology. Deep learning algorithms can provide accurate mappings given large numbers of labeled instances for training and validation. Generating adequate volume of quality labels has emerged as a critical barrier in computational pathology given the time and effort required from pathologists. In this paper we describe an approach for engaging crowds of medical students and pathologists that was used to produce a dataset of over 220,000 annotations of cell nuclei in breast cancers. We show how suggested annotations generated by a weak algorithm can improve the accuracy of annotations generated by non-experts and can yield useful data for training segmentation algorithms without laborious manual tracing. We systematically examine interrater agreement and describe modifications to the MaskRCNN model to improve cell mapping. We also describe a technique we call Decision Tree Approximation of Learned Embeddings (DTALE) that leverages nucleus segmentations and morphologic features to improve the transparency of nucleus classification models. The annotation data produced in this study are freely available for algorithm development and benchmarking at: https://sites.google.com/view/nucls.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge